Science Spotlight
Appending our knowledge of limb evolution: new insights from zebrafish mutants
By Prableen K. Chowdhary
Science Spotlight highlights one or more recent papers that are of broad interest to the zebrafish community and make a notable impact on their field. The IZFS board would like to encourage trainees who are interested in science communication to get involved in interviewing authors and writing pieces for this section in future issues of the News Splash. If you would like to participate in preparing a piece for the next newsletter, please contact Cecilia Moens (cmoens@fredhutch.org).
The Spring 2021 Science Spotlight article is written by Prableen K. Chowdhary. Prableen is a graduate student in the Brewster Lab at the University of Maryland, Baltimore County, where she studies how circadian rhythms are altered under hypoxia in zebrafish embryos. In this issue, Prableen chose to write about a recent paper in Cell from Matthew Harris’s lab: Hawkins et al., (2021) Latent potential to form limb-like structures in zebrafish.
https://www.sciencedirect.com/science/article/pii/S0092867421000039?via%3Dihub

While uncovering the enigmatic phenomena that underlie hundreds of millions of years of evolution may seem like a daunting task to some, recent advances in technology have made this more feasible than ever. Thankfully, some brave scientists are up for the challenge. Recent work at Harvard University has shed new light on one particular evolutionary puzzle of limb formation: how did fins evolve into limbs? Fossil-based evidence suggests that the common ancestor of teleosts and tetrapods had pectoral appendages containing a series of side-by-side long bones along the anterior-posterior axis, the posterior-most of which was segmented to create an end-on-end articulation of bones called the metapterygium. This end-on articulation is hypothesized to be the foundation for tetrapod limb formation (see schematic). The existing paradigm has been that this structure is not retained in teleost fishes, but researchers at Harvard University are challenging that. In their recent paper in Cell (PMID: 33545089), researchers M. Brent Hawkins, Katrin Henke, and Matthew Harris uncover a novel developmental program that governs the formation of limb-like structures in zebrafish. 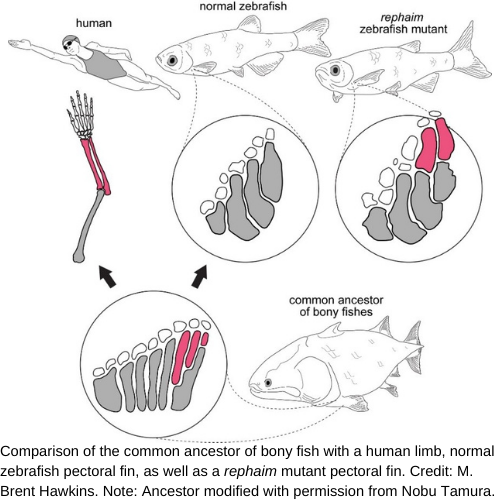
The pectoral fins of fish are homologous to tetrapod forelimbs. Since both fins and limbs are derived from locomotive organs in their common ancestor, these appendages allow for high mobility in their respective hosts. While the fin bud in teleosts gives rise a single row of long bones, the limb bud in tetrapods is segmented during development to create proximal (upper arm) and distal (forearm) bones connected by a joint in between. Despite the stark morphological contrast between fins and limbs, comparative approaches have identified shared developmental pathways responsible for the patterning of these appendages. Using a forward-genetics approach, the authors discovered a unique mutant, termed “rephaim,” that develops new skeletal elements along the pectoral fin proximal-distal axis in a limb-like manner (see schematic). “I actually went on my Facebook to ask who has good ideas for a mutant name,” Hawkins joked. “One of my friends – an old scholar at Harvard Divinity School – suggested rephaim because it refers to a ‘race of giants’ in the Old Testament that were said to have an extra finger on each hand and an extra toe on each foot.” Similar to long bones in tetrapods, skeletal staining revealed that the limb-like bone organization in rephaim mutants results from the segmentation of the long bones into proximal and distal regions separated by an “interzone.” Histological staining then revealed that these interzones give rise to joints and sites for muscle attachments in a manner that has not previously been reported in the pectoral fins of any other teleost.
Notably, this phenotype was mapped to gain-of-function mutations in waslb, which encodes a member of the WAS protein family of actin cytoskeleton signaling regulators previously implicated in cell migration, cell signaling, cell polarity, and transcriptional regulation. A second, phenotypically similar mutant was shown to be a gain-of-function mutation in vav2. The Vav family of protooncogenes encode RhoGEF proteins that have roles in signal transduction, cytoskeletal regulation, and cell motility. In the fin, waslb and vav2 gain-of-function mutations cause ectopic hox11 gene expression, and the authors show that hox11 genes are required for the formation of the extra skeletal elements in waslb mutants. How proteins that are primarily implicated in actin dynamics impact Hox expression remains an open question. “This ability for limb-like development in these fish was totally unexpected” says Hawkins. “When we found these genes waslb and vav2 that have had no previously known role in limb development, the next question really is: how do these genes interact with hox genes to cause this change in patterning? It’s challenging but exciting.”
Interestingly, however, Wasl conditional knockout mice also showed defects in skeletal formation, suggesting that this joint- and appendage-patterning pathway may be conserved in vertebrates. The zebrafish mutants show that even after millions of years of divergence between tetrapods and non-tetrapods, teleosts retain a “cryptic ancestral axis of growth,” almost akin to a memory for limb-forming potential.
About the authors: M. Brent Hawkins is a recent doctoral recipient from the Department of Organismic and Evolutionary Biology at Harvard University. In a pre-pandemic world, Hawkins enjoyed going to rock concerts and being around large groups of people. In current days he’s been fixing up a 20-year-old Buick Century sedan – a car that now deserves quite a bit of credit for getting him to the lab during a lockdown without public transportation. Matthew Harris is a professor at Harvard University and a researcher at Boston Children’s Hospital. His lab studies the genetic regulation of postembryonic development and its role in generation of form, physiology, and manifestation of disease. Nowadays Harris spends his personal time with his children and family.




